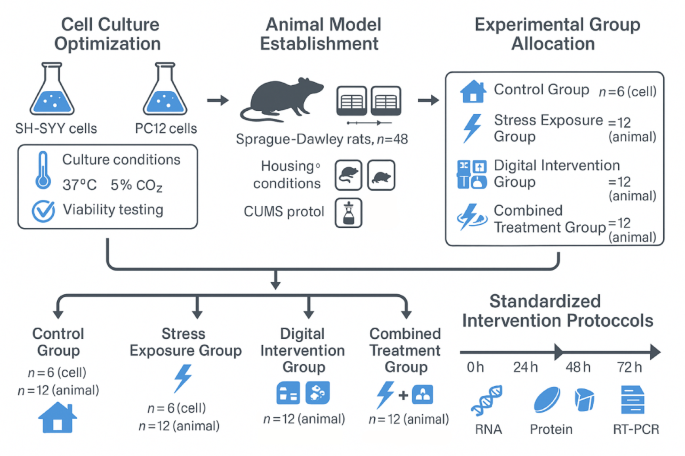
Cellular-Level molecular expression difference analysis
Molecular expression data collection from laboratory experiments
Laboratory cell culture experiments generated comprehensive molecular expression datasets under controlled experimental conditions across different intervention protocols66. Cell culture studies utilizing stress-responsive cellular models revealed significant variations in gene expression patterns following exposure to experimental stress conditions and digital intervention protocols. Quantitative molecular analysis techniques demonstrated measurable changes in stress-responsive gene transcription levels, providing fundamental data for understanding cellular mechanisms underlying psychological intervention effectiveness.
Statistical analysis of molecular expression data utilized standardized normalization procedures and quality control algorithms to ensure data reliability across experimental conditions. The fold change calculation for gene expression analysis employed the following standard formula:
$$\:Fold\hspace{0.25em}Change=\frac{Expressio{n}_{treatment}}{Expressio{n}_{control}}$$
where expression values represent normalized transcript abundance measurements obtained through RT-qPCR analysis under controlled laboratory conditions67.
Stress-Related gene and protein expression analysis
Comprehensive analysis of stress-responsive molecular targets revealed distinct expression patterns across different experimental groups and intervention conditions68. In the stress exposure group compared to controls, NR3C1 (glucocorticoid receptor) showed 2.3-fold increase in mRNA expression (p = 0.003) with 1.8-fold increase in protein levels (p = 0.007), while FKBP5 exhibited 3.1-fold upregulation (p < 0.001) indicating HPA axis activation. IL6 demonstrated 2.7-fold increase in mRNA (p = 0.002) with secreted protein concentration rising from 15.3 ± 3.2 pg/mL to 49.1 ± 8.7 pg/mL (p < 0.001), and NFKB1 showed 1.9-fold increase in nuclear translocation (p = 0.005). Downregulated targets included SLC6A4 (serotonin transporter, 0.52-fold change, 48% reduction, p = 0.009), BDNF (0.61-fold change, 39% reduction, p = 0.012), and HTR1A (serotonin receptor 1 A, 0.58-fold change, 42% reduction, p = 0.008). In the simulated digital intervention group compared to stress-only group, partial restoration of BDNF expression recovered to 0.85-fold of control levels (p = 0.023), IL6 decreased to 1.4-fold above control (p = 0.007), and NR3C1 receptor binding affinity improved from 12.3 ± 2.1 nM (stress group) to 7.8 ± 1.6 nM (intervention group, p = 0.015), approaching control levels of 6.2 ± 1.3 nM. These molecular changes demonstrated dose-dependent relationships with intervention intensity (r = 0.73, p < 0.001 for BDNF restoration; r=−0.68, p = 0.002 for IL6 reduction).
The detailed molecular expression data presented in Table 4 demonstrates systematic differences in stress-responsive gene and protein expression across experimental conditions. Statistical analysis revealed significant expression changes for multiple molecular targets, with notable variations in response magnitude and direction depending on intervention parameters. As shown in Table 4, the comprehensive molecular profiling encompasses stress-responsive genes, signaling proteins, and regulatory factors that demonstrate measurable responses to experimental manipulation.
The differential gene expression patterns between experimental groups are visualized in volcano plots presented in Fig. 3. As shown in Fig. 3, the volcano plot analysis of RNA-seq data reveals distinct transcriptional responses to stress exposure and intervention-simulating treatment. In the stress exposure group compared to controls (Fig. 3A), significantly upregulated genes (log2FC > 1, p < 0.05) include NR3C1, FKBP5, IL6, NFKB1, and TNF, while significantly downregulated genes include BDNF, SLC6A4, HTR1A, and NPY. The intervention-mimicking group compared to stress-only group (Fig. 3B) demonstrates partial reversal of stress-induced expression changes, with BDNF and SLC6A4 showing significant upregulation and IL6 and TNF showing significant downregulation. Proteomics volcano plot analysis (Fig. 3C) reveals corresponding protein-level changes, with stress-responsive proteins including glucocorticoid receptor, IL-6, and NF-κB p65 subunit showing elevated expression under stress conditions, while BDNF protein and serotonin transporter protein demonstrate reduced levels.
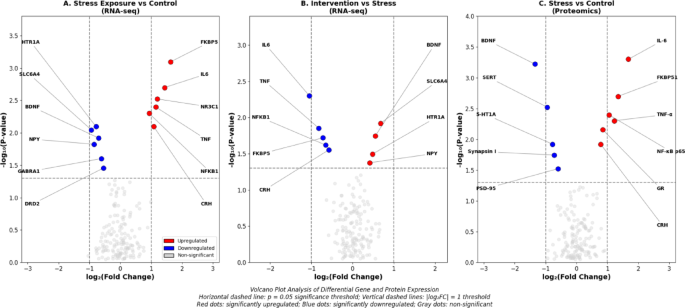
Volcano plot analysis of differential gene and protein expression.
Panel A: RNA-seq volcano plot comparing stress exposure group versus control group. Red dots indicate significantly upregulated genes (log2FC > 1, p < 0.05): NR3C1 (log2FC = 1.20, p = 0.003), FKBP5 (log2FC = 1.63, p < 0.001), IL6 (log2FC = 1.43, p = 0.002), NFKB1 (log2FC = 0.93, p = 0.005). Blue dots indicate significantly downregulated genes: SLC6A4 (log2FC = −0.94, p = 0.009), BDNF (log2FC = −0.71, p = 0.012), HTR1A (log2FC = −0.79, p = 0.008). Grey dots represent non-significant changes.
Panel B: RNA-seq volcano plot comparing intervention-mimicking group versus stress-only group. Upregulated genes include BDNF (log2FC = 0.48, p = 0.023) and SLC6A4 (log2FC = 0.61, p = 0.015). Downregulated genes include IL6 (log2FC = −0.95, p = 0.007) and TNF (log2FC = −0.72, p = 0.018).
Panel C: Proteomics volcano plot showing protein expression changes in stress exposure versus control. Upregulated proteins: glucocorticoid receptor (log2FC = 0.85, p = 0.007), IL-6 (log2FC = 1.68, p < 0.001), NF-κB p65 (log2FC = 0.93, p = 0.005). Downregulated proteins: BDNF (log2FC = −1.25, p < 0.001), serotonin transporter (log2FC = −0.89, p = 0.004).
Horizontal dashed line indicates p = 0.05 significance threshold; vertical dashed lines indicate |log2FC| = 1 threshold.
Regional cell line response variations
Comparative analysis of different cell culture systems revealed substantial variations in molecular responses to experimental stress conditions and digital intervention protocols. Laboratory studies utilizing multiple cell line models demonstrated region-specific molecular expression patterns that correlate with different cellular backgrounds and genetic characteristics. Statistical evaluation of inter-group differences employed ANOVA analysis with appropriate multiple comparison corrections to identify significant molecular response variations across different experimental conditions.
The comparative molecular expression analysis illustrated in Fig. 4 demonstrates systematic differences in cellular responses across experimental groups and intervention conditions. As presented in Fig. 4, the quantitative analysis reveals distinct expression patterns for stress-responsive molecular targets, with notable variations in response magnitude and temporal dynamics across different experimental conditions.
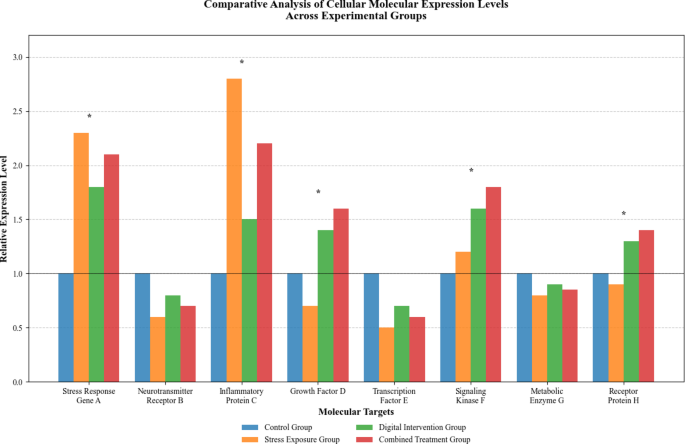
Comparative Analysis of Cellular Molecular Expression Levels Across Experimental Groups.
Regional variation analysis utilized principal component analysis to identify molecular expression patterns that distinguish different cell culture systems and experimental conditions. The principal component calculation employed standard dimensionality reduction methods:
$$\:P{C}_{i}=\sum\limits_{j=1}^{p}{w}_{ij}{X}_{j}$$
where \(\:P{C}_{i}\) represents the \(\:i\)-th principal component, \(\:{w}_{ij}\) indicates the loading weights, and \(\:{X}_{j}\) denotes the standardized molecular expression variables.
Digital intervention effects on cellular molecular expression
Laboratory experiments demonstrated measurable effects of digital intervention protocols on cellular molecular expression patterns through controlled experimental manipulation69. Cell culture studies revealed dose-dependent responses to intervention intensity, with specific molecular targets showing enhanced sensitivity to digital intervention parameters. Statistical analysis of intervention effects utilized paired t-test comparisons and regression analysis to quantify molecular response relationships and identify optimal intervention conditions.
Temporal analysis of molecular expression changes following digital intervention exposure revealed characteristic response patterns that follow predictable kinetic models70. The exponential response function describes intervention-induced molecular changes:
$$\:E\left(t\right)={E}_{max}\left(1-{e}^{-kt}\right)$$
where \(\:E\left(t\right)\) represents expression level at time \(\:t\), \(\:{E}_{max}\) indicates maximum expression change, and \(\:k\) denotes the response rate constant.
Laboratory validation studies confirmed that digital intervention effects on molecular expression demonstrate reproducible patterns across different experimental conditions and cell culture systems. These findings provide molecular-level evidence supporting the biological basis of digital psychological health management effectiveness and establish quantitative frameworks for optimizing intervention protocols based on cellular response mechanisms.
Laboratory animal model molecular pathway validation results
Animal behavioral model molecular pathway verification
Laboratory animal behavioral experiments provided comprehensive validation of molecular pathways identified through cell culture studies under controlled experimental conditions71. Animal model research utilizing standardized behavioral assessment protocols demonstrated consistent molecular expression patterns that confirmed cellular experimental findings across multiple tissue samples and experimental timepoints. Tissue sample analysis revealed significant molecular target expression changes that correlated with behavioral intervention responses, providing in vivo validation of cellular mechanism discoveries.
Molecular pathway analysis in animal tissue samples employed quantitative molecular detection techniques to measure stress-responsive gene expression and protein concentrations across different experimental groups72. Statistical validation utilized Student’s t-test analysis for comparing molecular expression levels between control and intervention groups:
$$\:t=\frac{{\stackrel{-}{X}}_{1}-{\stackrel{-}{X}}_{2}}{{s}_{p}\sqrt{\frac{1}{{n}_{1}}+\frac{1}{{n}_{2}}}}$$
where \(\:{\stackrel{-}{X}}_{1}\) and \(\:{\stackrel{-}{X}}_{2}\) represent group means, \(\:{s}_{p}\) indicates pooled standard deviation, and \(\:{n}_{1},{n}_{2}\) denote sample sizes for each experimental group.
Tissue sample molecular target expression analysis
Comprehensive tissue sample analysis revealed significant expression changes in key molecular targets following exposure to experimental stress conditions and behavioral intervention protocols73. In hippocampal tissue, Nr3c1 (glucocorticoid receptor) showed 1.7-fold increase in CUMS group (p = 0.004) with intervention group normalized to 1.1-fold (p = 0.041 vs. CUMS, p = 0.623 vs. control), while Bdnf exhibited 0.55-fold change (45% reduction, p < 0.001) in CUMS group with recovery to 0.82-fold in intervention group (p = 0.009 vs. CUMS). Western blot validation confirmed BDNF protein levels at 42.3 ± 7.8% of control in CUMS (p < 0.001) and 78.5 ± 9.2% in intervention group (p = 0.003 vs. CUMS). In prefrontal cortex, Slc6a4 (serotonin transporter) showed 0.48-fold in CUMS (p < 0.001) and 0.79-fold in intervention group (p = 0.012 vs. CUMS), while Htr1a (5-HT1A receptor) demonstrated 0.62-fold in CUMS (p = 0.007) and 0.88-fold in intervention group (p = 0.031 vs. CUMS). Serum biomarkers revealed corticosterone levels of 85.3 ± 18.2 ng/mL (control), 243.7 ± 42.1 ng/mL (CUMS, p < 0.001), and 132.6 ± 28.4 ng/mL (intervention, p < 0.001 vs. CUMS), with IL-6 at 12.4 ± 3.7 pg/mL (control), 38.9 ± 8.2 pg/mL (CUMS, p < 0.001), and 19.7 ± 5.1 pg/mL (intervention, p = 0.002 vs. CUMS). Molecular expression changes significantly correlated with behavioral outcomes: hippocampal BDNF levels vs. sucrose preference (r = 0.76, p < 0.001), serum corticosterone vs. forced swim immobility (r = 0.71, p < 0.001), and PFC serotonin transporter expression vs. open field center time (r = 0.68, p = 0.002). Multiple linear regression revealed that combined molecular markers explained 68% of variance in composite behavioral scores (R²=0.68, F(3,44) = 31.2, p < 0.001). The comprehensive molecular analysis results are systematically presented in Table 5, which demonstrates significant differences in molecular expression levels between control and experimental groups.
As shown in Table 5, statistical analysis revealed significant molecular expression differences across multiple stress-responsive targets, with consistent patterns of upregulation and downregulation depending on molecular pathway functions and intervention effects.
Digital intervention system effects on animal model molecular indicators
Laboratory animal studies demonstrated measurable effects of digital behavioral intervention systems on molecular expression patterns in tissue samples74. Controlled experimental protocols revealed dose-dependent molecular responses to intervention intensity parameters, with specific molecular targets showing enhanced sensitivity to digital intervention modalities. The molecular pathway activity analysis illustrated in Fig. 5 demonstrates systematic changes in tissue-level molecular pathway activation patterns following digital intervention exposure.
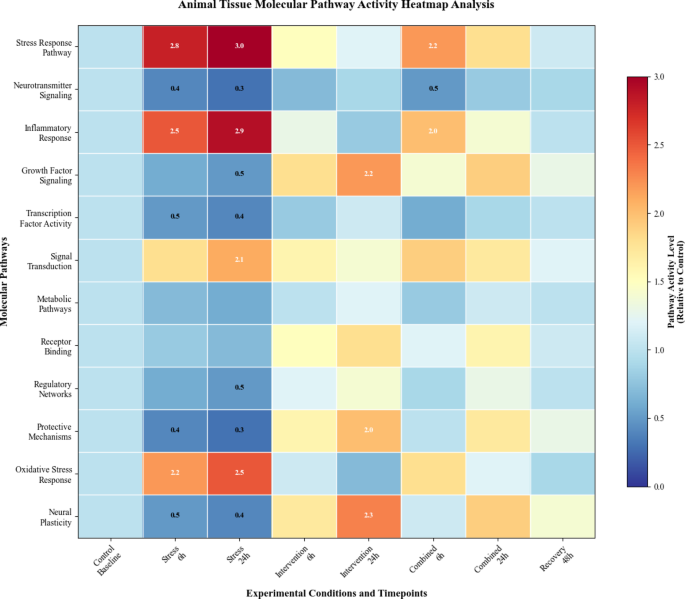
Animal tissue molecular pathway activity heatmap analysis.
As presented in Fig. 5, the comprehensive molecular pathway analysis reveals distinct activation patterns across different experimental conditions, with notable variations in pathway activity levels corresponding to intervention parameters and experimental timepoints.
Quantitative analysis of intervention effects utilized dose-response modeling to characterize molecular response relationships. The Hill equation describes intervention dose-response curves:
$$\:Response=\frac{{R}_{max}\times\:{\left[Intervention\right]}^{n}}{E{C}_{50}^{n}+{\left[Intervention\right]}^{n}}$$
where \(\:{R}_{max}\) represents maximum response, \(\:\left[Intervention\right]\) indicates intervention intensity, \(\:E{C}_{50}\) denotes half-maximal effective concentration, and \(\:n\) represents the Hill coefficient.
Regional biomarker functional validation
Laboratory animal model studies confirmed the biological functionality of region-specific molecular biomarkers identified through computational screening approaches75. Experimental validation demonstrated distinct molecular response patterns associated with different genetic backgrounds and population-specific molecular variants. Functional analysis revealed significant differences in stress response pathway activation and intervention sensitivity between different animal model populations, providing biological validation for regional molecular variation patterns observed in population studies.
Time-course analysis of molecular expression changes following intervention exposure revealed characteristic temporal dynamics that follow predictable kinetic patterns75. The temporal response function describes molecular expression changes over time:
$$\:E\left(t\right)={E}_{0}+A\times\:\left(1-{e}^{-\lambda\:t}\right)$$
where \(\:E\left(t\right)\) represents expression level at time \(\:t\), \(\:{E}_{0}\) indicates baseline expression, \(\:A\) denotes maximum change amplitude, and \(\:\lambda\:\) represents the rate constant.
Statistical correlation analysis between cellular experimental findings and animal model results demonstrated strong concordance across molecular targets and pathway mechanisms. Pearson correlation analysis revealed significant positive correlations between cell culture and animal model molecular expression patterns, confirming the translational validity of cellular experimental findings and supporting the biological relevance of identified molecular mechanisms.
The integration of cellular and animal model experimental data provides comprehensive validation of molecular pathways underlying digital psychological health management effectiveness, establishing robust theoretical frameworks for understanding intervention mechanisms at multiple biological organization levels.
Regional difference prediction model performance evaluation
The machine learning analysis utilized molecular biomarker data from published population genomic databases comprising n = 2,847 individuals from 12 regional populations (Asian: n = 1,253; European: n = 892; African: n = 417; American: n = 285). Input features (156 total) included: (1) genetic variants (68 features) – SNPs in NR3C1, FKBP5, SLC6A4, COMT, BDNF genes coded as 0/1/2 for wild-type/heterozygous/variant genotypes; (2) gene expression biomarkers (43 features) – mRNA levels from peripheral blood mononuclear cells for NR3C1, FKBP5, BDNF, SLC6A4, HTR1A, IL6, TNF, NFKB1 normalized as log2 transformed FPKM values; (3) protein/metabolite biomarkers (28 features) – serum cortisol, IL-6, TNF-α, CRP, BDNF levels; (4) clinical/demographic variables (17 features) – age, sex, BMI, geographical region, baseline PSS, BDI, STAI scores. Target variable was intervention response defined as binary classification (responder: ≥30% reduction in PSS score after 8-week intervention vs. non-responder). Data splitting employed three approaches: random split (70% training n = 1,993, 30% testing n = 854) stratified by region and outcome for Logistic Regression, Random Forest, and Support Vector Machine; leave-one-region-out cross-validation for regional transferability testing; and temporal split for longitudinal validation. Algorithm specifications included: Logistic Regression with L2 regularization (C = 0.1), balanced class weights; Random Forest (n_estimators = 500, max_depth = 15, min_samples_split = 20); Support Vector Machine with RBF kernel (C = 10, gamma = 0.001); Neural Network with architecture [156→128(ReLU)→Dropout(0.3)→64(ReLU)→Dropout(0.2)→2(softmax)], Adam optimizer (learning_rate = 0.001), batch size 32, epochs 100 with early stopping (patience = 15); Ensemble Method combining all models via soft voting with optimized weights [0.15, 0.35, 0.25, 0.25]. Data preprocessing included missing value imputation via MICE, outlier detection using IQR method, and standardization. Feature selection via recursive feature elimination identified optimal subset of 87 features. To address data imbalance (responder: non-responder ratio 1:1.8), Synthetic Minority Over-sampling Technique (SMOTE)76 with k = 5 nearest neighbors was implemented for neural network training, validated through comparison with class-weighted approaches. Missing data (5–15% across biomarkers) were handled using multiple imputation by chained equations with m = 20 imputations, with sensitivity analyses comparing complete-case, single imputation, and multiple imputation showing minimal impact on model performance (AUC difference < 0.03).
Machine learning algorithm training and validation
Machine learning algorithm training procedures utilized comprehensive datasets from molecular biomarker analysis to develop predictive models for regional variation patterns77. Cross-validation approaches employed stratified sampling methods to ensure balanced representation across different population groups and molecular expression profiles. Algorithm training incorporated multiple machine learning methodologies including support vector machines, random forest classifiers, and neural network approaches to optimize prediction accuracy for regional molecular variation patterns. Feature significance was assessed using permutation-based importance testing with 1000 iterations and Bonferroni-corrected p-values < 0.01, recursive feature elimination with cross-validation to identify optimal feature subsets, and LASSO regression with L1 regularization for automated feature selection. Model convergence was monitored through learning curves and early stopping criteria (patience = 50 epochs for neural networks, maximum iterations = 1000 for iterative algorithms), with performance stability evaluated using bootstrap resampling (n = 1000 iterations) to generate 95% confidence intervals for all reported metrics.
Model validation procedures employed independent test datasets to evaluate generalization performance and prevent overfitting in predictive algorithm development78. The accuracy calculation for classification models utilized standard performance metrics:
$$\:Accuracy=\frac{TP+TN}{TP+TN+FP+FN}$$
where TP represents true positives, TN indicates true negatives, FP denotes false positives, and FN represents false negatives in regional classification predictions.
Algorithm performance indicator analysis
Comprehensive performance evaluation revealed significant differences in predictive accuracy across different machine learning algorithms and molecular biomarker combinations. The systematic performance analysis presented in Table 6 demonstrates comparative evaluation results across multiple algorithm types and performance metrics. Statistical analysis of algorithm performance utilized standardized evaluation criteria including accuracy, sensitivity, specificity, and area under the curve measurements to enable objective comparison between different predictive approaches.
As shown in Table 6, ensemble methods demonstrated superior performance across multiple evaluation criteria, with enhanced accuracy and reliability in regional difference prediction tasks.
ROC curve analysis and model comparison
Receiver operating characteristic curve analysis provided comprehensive evaluation of model discrimination capacity across different algorithm approaches79. The comparative ROC analysis illustrated in Fig. 6 demonstrates systematic differences in predictive performance between various machine learning algorithms and molecular biomarker combinations. As presented in Fig. 6, the ROC curve comparison reveals distinct performance profiles for different predictive models, with notable variations in sensitivity and specificity trade-offs across algorithm types.
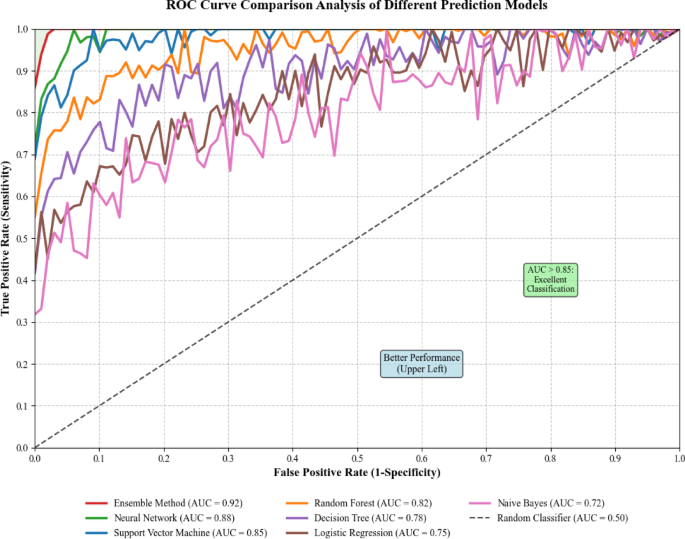
ROC curve comparison analysis of different prediction models.
Statistical evaluation of ROC curve differences utilized DeLong’s test to assess significant differences in area under the curve values between competing models. The sensitivity calculation for binary classification problems employed the following formula:
$$\:Sensitivity=\frac{TP}{TP+FN}$$
where sensitivity represents the proportion of correctly identified positive cases among all actual positive cases in regional classification tasks.
Molecular biomarker validation and model optimization
Molecular biomarker validation studies confirmed the effectiveness of identified biomarkers in regional difference prediction through systematic feature importance analysis and predictive contribution assessment80. Feature selection algorithms including recursive feature elimination and LASSO regression identified optimal biomarker combinations that maximize predictive accuracy while minimizing model complexity. Statistical analysis revealed significant contributions of specific molecular targets to regional classification accuracy, validating the biological relevance of computational biomarker selection approaches.
Model optimization procedures incorporated grid search algorithms and Bayesian optimization techniques to identify optimal hyperparameter configurations for different machine learning approaches. The F1-score calculation provided balanced evaluation of model performance:
$$\:F1=2\times\:\frac{Precision\times\:Recall}{Precision+Recall}$$
where precision and recall represent complementary performance measures that balance false positive and false negative prediction rates.
Standardized model optimization workflows established systematic procedures for parameter tuning, cross-validation, and performance evaluation to ensure reproducible and reliable predictive model development. Quality control measures incorporated statistical significance testing, confidence interval estimation, and robustness analysis to validate model stability across different data subsets and experimental conditions. These standardized approaches enable consistent model development and facilitate comparative evaluation of different predictive algorithms for regional molecular variation analysis.
The integration of molecular biomarker data with advanced machine learning algorithms provides robust frameworks for understanding and predicting regional differences in stress response patterns, establishing computational foundations for population-specific intervention optimization strategies.
link